Essential Information on Enzyme Activities Required for Effective Cell Isolation
Part A: Overview of Current Knowledge and State of the Art
| Overview |
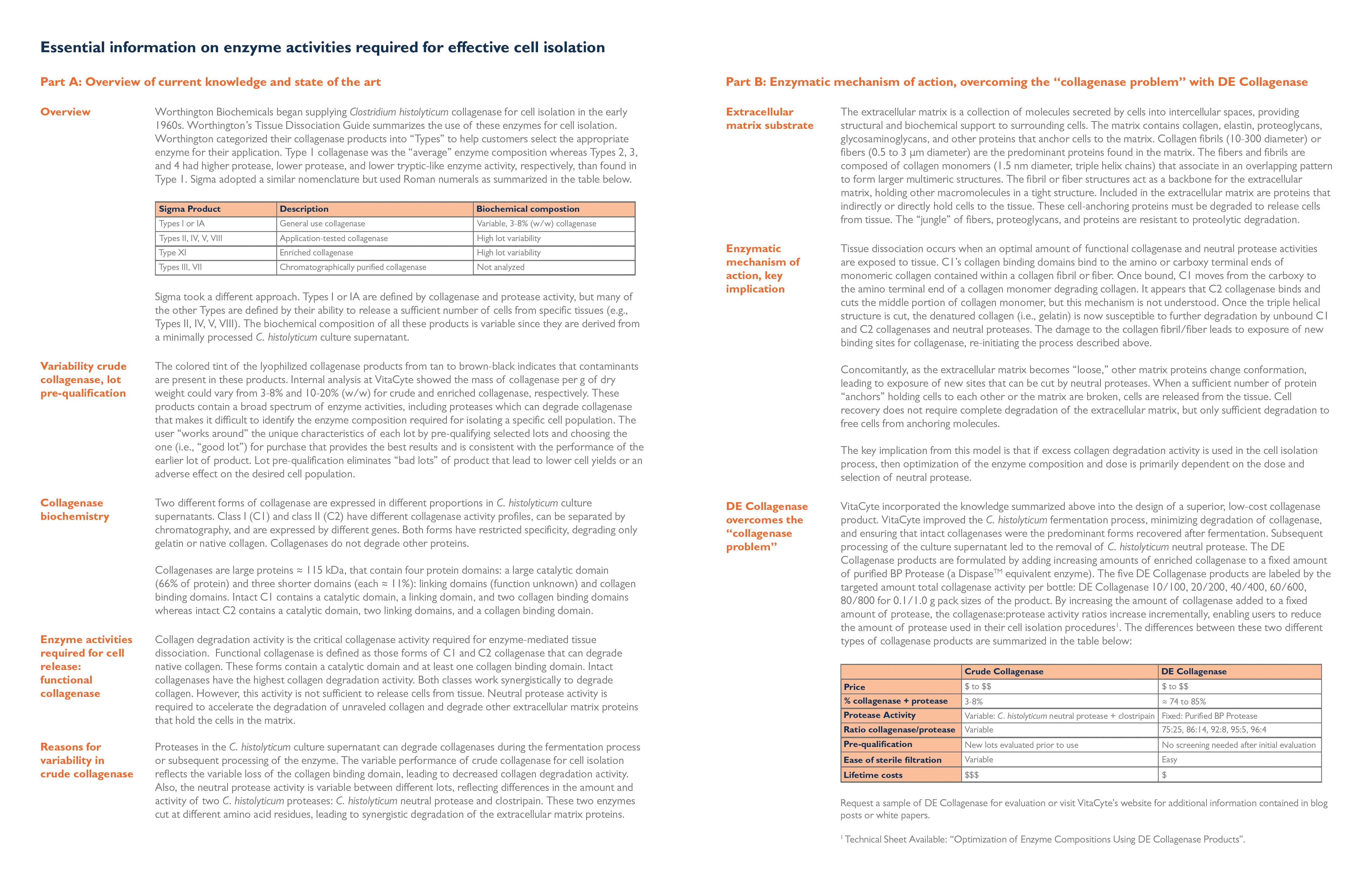
Worthington Biochemicals began supplying Clostridium histolyticum collagenase for cell isolation in the early 1960s. Worthington’s Tissue Dissociation Guide summarizes the use of these enzymes for cell isolation. They categorized their collagenase products into “Types” to help customers select the appropriate enzyme for their application. Type 1 collagenase was the “average” enzyme composition whereas Types 2, 3, and 4 had higher protease, lower protease, and lower tryptic-like enzyme activity, respectively, than found in Type 1. Sigma adopted a similar nomenclature but used Roman numerals as summarized in the table below:
Sigma took a different approach. Types I or IA are defined by collagenase and protease activity, but many of the other Types are defined by their ability to release a sufficient number of cells from specific tissues (e.g., Types II, IV, V, VIII). The biochemical composition of these products are variable since they are derived from a minimally processed C. histolyticum culture supernatant. |
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Variability of Crude Collagenase, Lot Pre-Qualification |
The brown color of the lyophilized collagenase products indicates the presence of contaminants. Analysis indicates only 3-8% of the brown powder is active enzymes and 10-20% (w/w) for crude and enriched collagenase, respectively. These products contain a broad spectrum of enzyme activities including proteases which can degrade collagenase. This complexity makes it difficult to identify the optimal enzyme composition required for isolating a specific cell population. The user “works around” the unique characteristics of each lot by pre-qualifying selected lots and choosing the one (i.e., “good lot”) for purchase that provides the best results and is consistent with the performance of the earlier lot of product. Lot pre-qualification eliminates “bad lots” of product that lead to lower cell yields or an adverse effect on the desired cell population. | |||||||||||||||
| Collagenase Biochemistry | Two different forms of collagenase are expressed in different proportions in C. histolyticum culture supernatants. Class I (C1) and class II (C2) have different collagenase activity profiles, can be separated by chromatography, and are expressed by different genes. Both forms have restricted specificity, degrading only gelatin or native collagen. Collagenases do not degrade other proteins. Collagenases are large proteins ≈ 115 kDa, that contain four protein domains: a large catalytic domain (66% of protein) and three shorter domains (each ≈ 11%): linking domains (function unknown) and collagen binding domains. Intact C1 contains a catalytic domain, a linking domain, and two collagen binding domains whereas intact C2 contains a catalytic domain, two linking domains, and a collagen binding domain. |
|||||||||||||||
| Enzyme Activities Required for Cell Release: Functional Collagenase |
Collagen degradation activity is the critical collagenase activity required for enzyme-mediated tissue dissociation. Functional collagenase is defined as those forms of C1 and C2 collagenase that can degrade native collagen. These forms contain a catalytic domain and at least one collagen binding domain. Intact collagenases have the highest collagen degradation activity. Both classes work synergistically to degrade collagen. However, this activity is not sufficient to release cells from tissue. Neutral protease activity is required to accelerate the degradation of unraveled collagen and degrade other extracellular matrix proteins that hold the cells in the matrix. | |||||||||||||||
| Reasons for Variability in Crude Collagenase | Proteases in the C. histolyticum culture supernatant can degrade collagenases during the fermentation process or subsequent processing of the enzyme. The variable performance of crude collagenase for cell isolation reflects the variable loss of the collagen binding domain, leading to decreased collagen degradation activity. Also, the neutral protease activity is variable between different lots, reflecting differences in the amount and activity of two C. histolyticum proteases: C. histolyticum neutral protease and clostripain. These two enzymes cut at different amino acid residues, leading to synergistic degradation of the extracellular matrix proteins. |
Part B: Enzymatic Mechanism of Action, Overcoming the “Collagenase Problem” with DE Collagenase
| Extracellular Matrix Substrate | The extracellular matrix is a collection of molecules secreted by cells into intercellular spaces providing structural and biochemical support to surrounding cells. The matrix contains collagen, elastin, proteoglycans, glycosaminoglycans, and other proteins that anchor cells to the matrix. Collagen fibrils (10-300 diameter) or fibers (0.5 to 3 μm diameter) are the predominant proteins found in the matrix. The fibers and fibrils are composed of collagen monomers (1.5 nm diameter, triple helix chains) that associate in an overlapping pattern to form larger multimeric structures. The fibril or fiber structures act as a backbone for the extracellular matrix, holding other macromolecules in a tight structure. Included in the extracellular matrix are proteins that indirectly or directly hold cells to the tissue. These cell-anchoring proteins must be degraded to release cells from tissue. The “jungle” of fibers, proteoglycans, and proteins are resistant to proteolytic degradation. | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Enzymatic Mechanism of Action, Key Implication | Tissue dissociation occurs when an optimal amount of functional collagenase and neutral protease activities are exposed to tissue. C1’s collagen binding domains bind to the amino or carboxy terminal ends of monomeric collagen contained within a collagen fibril or fiber. Once bound, C1 moves from the carboxy to the amino terminal end of a collagen monomer degrading collagen. It appears that C2 collagenase binds and cuts the middle portion of a collagen monomer, but this mechanism is not understood. Once the triple helical structure is cut, the denatured collagen (i.e., gelatin) is now susceptible to further degradation by unbound C1 and C2 collagenases and neutral proteases. The damage to the collagen fibril/fiber leads to exposure of new binding sites for collagenase, re-initiating the process described above. Concomitantly, as the extracellular matrix becomes “loose,” other matrix proteins change conformation leading to exposure of new sites that can be cut by neutral proteases. When a sufficient number of protein “anchors” holding cells to each other or the matrix are broken, cells are released from the tissue. Cell recovery does not require complete degradation of the extracellular matrix, but only sufficient degradation to free cells from anchoring molecules. The key implication from this model is that if excess collagen degradation activity is used in the cell isolation process, then optimization of the enzyme composition and dose is primarily dependent on the dose and selection of neutral protease. |
||||||||||||||||||||||||
| DE Collagenase Overcomes the “Collagenase Problem” |
VitaCyte incorporated the knowledge summarized above into the design of a superior, low-cost collagenase product. VitaCyte improved the C. histolyticum fermentation process, minimizing degradation of collagenase, and ensuring that intact collagenases were the predominant forms recovered after fermentation. Subsequent processing of the culture supernatant led to the removal of C. histolyticum neutral protease. The DE Collagenase products are formulated by adding increasing amounts of enriched collagenase to a fixed amount of purified BP Protease (a Dispase™ equivalent enzyme). The five DE Collagenase products are labeled by the targeted amount of total collagenase activity per bottle: DE Collagenase 10/100, 20/200, 40/400, 60/600, 80/800 for 0.1/1.0 g pack sizes of the product. By increasing the amount of collagenase added to a fixed amount of protease, the collagenase:protease activity ratios increase incrementally, enabling users to reduce the amount of protease used in their cell isolation procedures(1). The differences between these two different types of collagenase products are summarized in the table below:
|
1 Technical Sheet Available: “Optimization of Enzyme Compositions Using DE Collagenase Products”.