
Collagenase Quality
For many years, experts in cell isolation have stated the importance of collagenase quality for successful cell isolation.1,2 What does collagenase quality mean? Many investigators have found it difficult to correlate the enzymatic analysis data summarized on a Certificate of Analysis to the performance of a specific lot of crude or enriched collagenase to recover cells. This led the majority of users to rely on lot pre-qualification to select the best collagenase lot for cell isolation. This reliance on lot pre-qualification is likely responsible for the assumption that collagenase quality is important for cell isolation since the quality was determined by the end user.
Updated Knowledge
Today, collagenase “quality” can be explained by knowledge of biochemical characteristics of Clostridium histolyticum collagenase, specifically its ability to efficiently degrade native collagen.3 We know now that purified collagenase alone is insufficient to release cells from tissue. 4-7 C. histolyticum collagenases require neutral proteases to degrade other extracellular matrix proteins that hold cells to tissue and to accelerate the degradation of collagen.
To understand the mechanism of how enzymes release cells from tissue, we must consider essential features of the extracellular matrix, and biochemical characteristics of the collagenase required for cell release. The summary below provides a brief overview of the enzymes and tissue substrates involved in the isolation of primary cells from tissue. For additional information, request a copy of VitaCyte’s White Paper “Lot qualification: A common but costly pathway to select collagenase products for cell isolation. What are the known unknowns when using collagenase products?”
Extracellular Matrix
The extracellular matrix is a collection of molecules secreted by cells into the intercellular spaces, providing structural and biochemical support to surrounding cells. The matrix contains collagen, elastin, proteoglycans, glycosoaminoglycans, and other proteins that anchor cells to the matrix.8 Collagen fibrils (10-300 diameter) or fibers (0.5 to 3 μm diameter) are the predominant proteins found in the matrix.9 The fibers and fibrils are composed of collagen monomers (1.5 nm diameter) that associate in an overlapping pattern to form larger multimeric structures. The fibril or fiber structures act as backbone for the extracellular matrix, holding other macromolecules in a tight structure, resistant to proteolysis. Included in the extracellular matrix are proteins that indirectly or directly hold cells to the tissue. These cell-anchoring proteins must be degraded to release cells from tissue.
Characteristics of Collagenase
By definition, collagenase is any enzyme that cuts the triple helical protein chains characteristic of all collagen molecules. C. histolyticum collagenase is used for cell isolation because of its broad specificity to cut all forms of mammalian collagen. There are two different classes of C. histolyticum collagenase: class I (C1) and class II (C2) collagenase, identified by their difference in gelatinase and peptidase activities. C1 has strong gelatinase activity (measured by degradation of gelatin or denatured collagen) but low peptidase activity (measured by degradation of Pz peptide or FALGPA peptide). The converse is true for C2 collagenase.3 As will be noted below, these two classes work cooperatively to degrade native collagen.
Each class of collagenase is expressed as a separate gene and contains about 1000 amino acids, subdivided into 4 protein domains of ≈ 670, 110, 110, and 110 amino acids.10 The largest domain is the catalytic domain found at the amino terminal end of the protein followed by linking (function unknown) domain(s) and collagen binding domain(s) (CBDs) at the carboxy terminal end of the molecule. Each form is identified by its class and molecular weight expressed in kilodaltons (kDa). Intact C1 has a catalytic domain, one linking domain and two CBDs (C1116 kDa), whereas intact C2 has a catalytic domain, two linking domains and one CBD (C2114 kDa). The truncated form of C1 (C1100 kDa) has a catalytic domain, one linking domain and one CBD. It is generated after proteolysis of a short segment linking the two CBDs together, removing the terminal CBD from the molecule. This can occur if the bacterial culture supernatant has high protease activity or if the product is “mishandled” after fermentation.11
The figure below is the crystal structure of each domain found in intact C1 with a schematic outline of the protein domain structure below.12-14 If the structure was determined for the entire enzyme, all of the protein domains would be linked together.
At this time, the function of the linking domains is unknown. The assignment of other domains as catalytic or CBD is determined by their ability to function independently of any other protein domains. The 3 dimensional structure of intact C1 is shown below with a schematic representation of the protein domain structure.
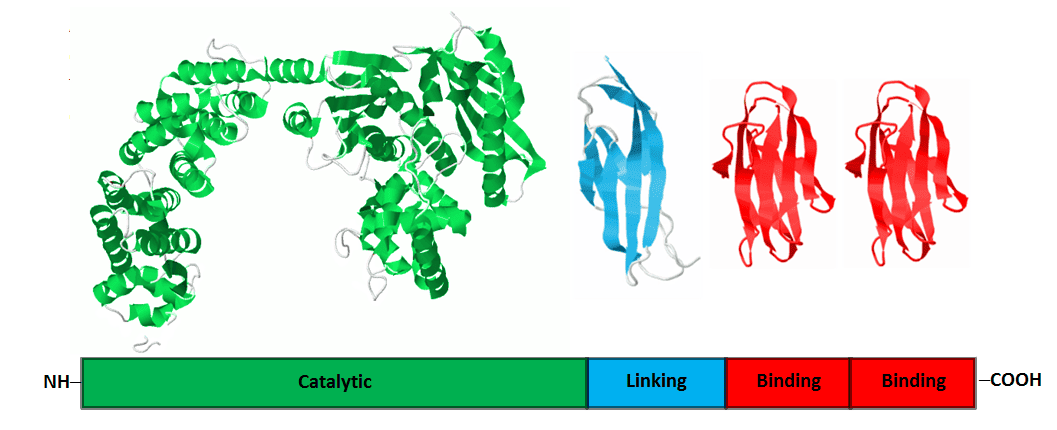
Three forms of collagenase, containing a catalytic domain and at least one CBD, are effective in degrading native collagen. These three enzymes ― C1116 kDa, C2114 kDa, and C1100 kDa ― are termed “functional collagenase” since they possess the unique ability to degrade the fundamental unit of all collagen forms: a tightly wound, triple helical protein chain. Subsequent studies showed purified intact C1 with two CBDs had 7-10 fold higher collagen degradation activity than purified intact C2 or purified truncated C1.15 Collagenase-protease enzyme mixtures containing primarily intact C1 and C2 with an optimal dose of protease are the highest quality and most efficient collagenase enzyme products because of a high specific collagen degradation activity (CDA U/mg protein).3 This statement is supported by results comparing the efficiency of purified collagenase from VitaCyte or from another supplier to recover islets from human pancreas. The use of VitaCyte’s product that contained primarily C1116 kDa, C2114 kDa and neutral protease gave significantly higher islet yields and shorter digestion times than those isolations performed using another supplier’s enzymes that contained primarily C1100 kDa and C2114 kDa and neutral protease.16
“Non-functional collagenases” are defined as collagenases with a functional catalytic domain, but no CBDs. The inability of these enzymes to bind native collagen means they are active only on those substrates that have a functional catalytic domain. These substrates include collagenase-specific peptide substrates (Pz or FALGPA peptide) or gelatin. Gelatin is a non-specific substrate since other proteases also degrade this protein.
Functional collagenase also has gelatinase and peptidase activities, so gelatin, Pz and FALGPA peptide substrates cannot distinguish functional from non-functional collagenase.
Hypothetical Mechanism of Tissue Dissociation
When tissue is exposed to an optimal amount of functional collagenase and neutral protease activities, C1 collagenase starts collagen degradation by binding to the amino or carboxy terminal ends of monomeric collagen contained within a collagen fibril or fiber. It appears that C2 collagenase binds and cuts the middle portion of collagen monomer.17 Once bound, the catalytic domain can cut collagen’s triple helical structure. Each catalytic domain can cut many collagen molecules tightly packed in collagen fibrils or fibers. Once the triple helical structure is cut, the three strands unravel, leading to the formation of denatured collagen (i.e., gelatin). These portions of denatured collagen are now susceptible to further degradation by unbound C1 and C2 collagenases and neutral proteases. Collagenolytic and proteolytic activity releases sections of collagen from the fiber or fibril, leading to exposure of new binding sites for collagenase, re-initiating the process described above.
Concomitantly, as the extracellular matrix becomes “loose”, other matrix proteins change conformation, leading to exposure of new sites that can be cut by neutral protease. When a sufficient number of protein “anchors” holding cells to each other or to the matrix are broken, cells are released from the tissue. Cell recovery does not require complete degradation of the extracellular matrix, but only sufficient degradation to free cells from anchoring molecules.
Implications from New Knowledge
The adage that collagenase quality is important for successful cell isolation is only partially correct. Suboptimal amounts of functional collagenase activity leads to incomplete degradation of native collagen, decreasing the efficiency of protease to effectively degrade cell anchoring proteins that hold cells to the extracellular matrix. This results in suboptimal recovery of cells from tissue. However, if the collagenase products which contain primarily intact C1 and C2 and are used in excess, then the focus should be on selection and dose of neutral protease activity. Neutral protease activity determines the speed of digestion. Excess neutral protease activity damages cells, leading to low viability and cell yields. In contrast, excess collagenase is unlikely to have an adverse effect on the isolated cells for two reasons. First, degradation of native collagen only occurs when the enzyme is physically bound to a specific site on native collagen. Collagen is not an integral cell protein, but is present only in the extracellular matrix. Second, unbound functional or non-functional collagenase has narrow selectivity, degrading primarily denatured collagen (gelatin). It does not degrade other proteins.
Finding a Source of Low Cost Collagenase Containing Primarily Intact C1 and C2
Assuming that the highest quality collagenase contains primarily intact C1 and C2, your focus should be on finding a source of these defined and characterized enzymes. Biochemical analysis of Worthington Type I and Sigma Type XI collagenase lots has been performed by VitaCyte and shown to have variability in their amount of intact C1.11 Since the amount of intact C1 varies between lots, the efficiency of these collagenase lots to degrade native collagen also varies. Sigma and Worthington report collagen degradation activity per mg dry weight product using the Mandl assay (measures the degradation of insoluble collagen fibers). However, we have found recently that this assay is imprecise and unable to distinguish between intact and truncated C1 (unpublished data). In contrast, VitaCyte’s collagen degradation activity assay uses soluble fluorescein isothiocyanate labeled calf skin fibrils as substrate, with enzyme activity kinetically measured by fluorescent microplate reader. This assay has been shown to readily detect differences between specific collagen degradation activities of intact and truncated C1.15
VitaCyte’s development of a superior collagen degradation activity assay reflects its commitment to manufacture the highest quality collagenase and protease enzymes to improve the productivity of scientists isolating cells from tissue. VitaCyte recently introduced a new, low cost enriched collagenase product line (DE Collagenase) that contains primarily intact C1 and C2 collagenase. Each product is identified by the targeted amount of FALGPA activity per bottle. The five DE Collagenase products are offered as two pack sizes (100 mg or 1 g) containing increasing amounts of high quality collagenase (DE Collagenase 10/100, 20/200, 40/400, 60/600, 80/800) added to a fixed amount of purified neutral protease (BP Protease). For example the DE 80 (100 mg) or DE 800 (1 g) will have an 8 fold higher amount of collagenase activity than the DE 10 (100 mg) or DE 100 (1 g). The DE Collagenase formulations enable users to determine the impact of protease activity in their cell isolation procedure, since these enzyme mixtures contain excess collagenase activity.
References
- Berry MN, Edwards AM, Barritt GJ. Isolated hepatocytes: prepartion, properties and applications. New York: Elsevier; 1991.
- Scharp DW. Commentary. Cell Transplantation 1993; 2: 299.
- RC McCarthy RC AB, ML, FE Dwulet. Tissue dissociation enzymes for isolating human islets for transplantation: Factors to consider in setting enzyme acceptance criteria. Transplantation 2011; 91: 137-45.
- Hatton MW, Berry LR, Krestynski F, Sweeney GD, Regoeczi E. The role of proteolytic enzymes derived from crude bacterial collagenase in the liberation of hepatocytes from rat liver. Identification of two cell-liberating mechanisms. Eur J Biochem 1983; 137(1-2): 311-8.
- Hefley TJ, Stern PH, Brand JS. Enzymatic isolation of cells from neonatal calvaria using two purified enzymes from Clostridium histolyticum. Experimental Cell Research 1983; 149(1): 227-36.
- Suggs W, Van Wart H, Sharefkin JB. Enzymatic harvesting of adult human saphenous vein endothelial cells: use of a chemically defined combination of two purified enzymes to attain viable cell yields equal to those attained by crude bacterial collagenase preparations. Journal of Vascular Surgery 1992; 15(1): 205-13.
- Wolters GH, Vos-Scheperkeuter GH, van Deijnen JH, van Schilfgaarde R. An analysis of the role of collagenase and protease in the enzymatic dissociation of the rat pancreas for islet isolation. Diabetologia 1992; 35(8): 735-42.
- Frantz C, Stewart KM, Weaver VM. The extracellular matrix at a glance. Journal of Cell Science 2010; 123(Pt 24): 4195-200.
- Birk DE, Bruckner P. Collagen suprastructures. Topic Current Chemistry 2005; 247: 185-205.
- Matsushita O, Okabe A. Clostridial hydrolytic enzymes degrading extracellular components. Toxicon 2001; 39(11): 1769-80.
- VitaCyte. Lot qualification: a common but costly pathway to select collagenase products for cell isolation. Whar are the know unknowns when using collagenase products? 2017 (accessed 30 April 2017 2017).
- PDB R. Image from the RCSB PDB (www.rcsb.org) of PDB ID 1NQJ (Wilson JJ, Matsushita O, Okabe A, et al.) (2003) A bacterial collagen-binding domain with novel calcium-binding motif controls domain orientation Embo J 22:1743-52). 2003 (accessed 22 February 2017 2017).
- PDB R. Image from RCSB PDB (www.rcsb.org) of PDB ID 4TN9 (Bauer R, Janowska K, Taylor K, et al.) 2015) Structures of three polycystiuc kidney disease-like domains from Clostridium histolyticum collagenase ColG and ColH. Acta Crystallogra Sci Sect D 71:565-77). 2015 (accessed 22 February 2017 2017).
- PDB R. Image from the RCSB PDB (www.rcsb.org) of PDB ID 4ARE (Eckhard U, Schonauer E, Brandstetter H) (2013) Structural basis for activity regulation and substrate reference of Clostridium collagenases G, H, and T. J Biol Chem 288:20184). 2013 (accessed 22 February 2017 2017).
- McCarthy RC, Spurlin B, Wright MJ, et al. Development and characterization of a collagen degradation assay to assess purified collagenase used in islet isolation. Transplantation Proceedings 2008; 40(2): 339-42.
- Balamurugan AN, Breite AG, Anazawa T, et al. Successful human islet isolation and transplantation indicating the importance of class 1 collagenase and collagen degradation activity assay. Transplantation 2010; 89(8): 954-61.
- Watanabe-Nakayama T, Itami M, Kodera N, Ando T, Konno H. High-speed atomic force microscopy reveals strongly polarized movement of clostridial collagenase along collagen fibrils. Scientific Reports 2016; 6: 28975.
